
HOMER
Software for motif discovery and next-gen sequencing analysis
Initial Hi-C data processing with HOMER
Hi-C and variant technologies use paired-end sequencing to
assign regions of the genome to the same physical
location. After sequencing, you should have two files
(or many sets of 2 files) for the first and second reads
from each pair. Unlike paired end sequencing for other
techniques like genomic resequencing or RNA-Seq, where you
might expect the 2nd read to be located within the general
vicinity of the first read, read-pairs from Hi-C should be
processed independently (e.g. do not use Tophat, bowtie, bwa
or any short read alignment software in "paired-end" mode -
each read should be mapped independently!).Once Hi-C reads have been mapped, the makeTagDirectory program is used to process the reads. Hi-C has many more sources of noise than expected, so this step is particularly important, and may need to be revisited and tweaked. For this reason, we recommend running the command twice - once to complete the lengthy process of merging all the paired end reads into an "unprocessed" tag directory, then a second time on copies of the first tag directory using different parameters. For example:
makeTagDirectory HiC-Unprocessed
reads1-1.sam,reads1-2.sam reads2-1.sam,reads2-2.sam -tbp
1
Then experiment with different processing parameters (remember to include "-update" - this tells the program not to look for new data files but reprocess the existing tag directory)
This is a more efficient way to approach Hi-C processing if you're not sure what the data looks like yet.
Then experiment with different processing parameters (remember to include "-update" - this tells the program not to look for new data files but reprocess the existing tag directory)
cp -r HiC-Unprocessed/ HiC-HindIIIonly/
makeTagDirectory HiC-HindIIIonly -update -restrictionSite AAGCTT -both -genome hg19 -removePEbg
cp -r HiC-Unprocessed/ HiC-noSelfLigation/
makeTagDirectory HiC-noSelfLigation -update -restrictionSite AAGCTT -removeSelfLigation -removePEbg -genome hg19
makeTagDirectory HiC-HindIIIonly -update -restrictionSite AAGCTT -both -genome hg19 -removePEbg
cp -r HiC-Unprocessed/ HiC-noSelfLigation/
makeTagDirectory HiC-noSelfLigation -update -restrictionSite AAGCTT -removeSelfLigation -removePEbg -genome hg19
This is a more efficient way to approach Hi-C processing if you're not sure what the data looks like yet.
Creating a Paired-End Tag Directory with HOMER
Use makeTagDirectory to
process paired-end sequencing into a tag directory for use
with HOMER programs. For Hi-C, you want to map the
reads independently. This is different than most
"paired-end" alignments (such as for RNA-Seq) that
constrain the relative position of the fragments.
For Hi-C these assumptions do not hold. I'd
recommend mapping the same way you would ChIP-Seq data,
allowing only unique genomic matches. You may also want to
check for restriction sites in your reads - sequence on
either side of the restriction site might map to different
locations, so it might be wise to trim the sequence after
the restriction site to help with mapping (i.e. use "homerTools trim -3 AAGCTT reads.fq" to trim the
reads before mapping)
To create a paired-end tag directory, you must run makeTagDirectory by entering the two halves of the mapped reads separated by a comma, without spaces. For example:
HOMER matches reads between alignment files by matching their names, but to work with the original bowtie output files for Illumina sequencing (which often contains a 0 or 1 at the end to delineate the read source), it will need to remove the last character before trying to match. In this case, add "-illuminaPE" to the end of the command. If you download reads from the SRA they often have the same name for each paired end read - in these cases you don't want to remove the last character, so do NOT add "-illuminaPE" to the end of the command in that case.
Another typical option to add for Hi-C data is "-tbp 1", which will only consider read pairs with the exact same ends once. Given the search space in Hi-C, you would never expect to see reads-pairs start from the exact same places twice in the data (at least in the case of sonication). If you do, they are likely clonal and a result of over-sequencing your library or some other artifact.
On a side note, the HOMER tag directory created for paired-end reads is very similar to the directories created when running makeTagDirectory for single-end reads. The paired end reads are stored in *.tags.tsv files (extended format to include the pair), only each read will get 2 entries in the appropriate *.tags.tsv so that the reads can be indexed on each chromosome in one file. A flag is also set in the tagInfo.txt file to inform HOMER to treat the experiment as a paired-end file. Many of the programs in HOMER, such as makeUCSCfile, will treat paired-end tag directories as "single-end" directories to create output. This can be useful if you want to visualize read coverage in the UCSC Genome Browser or want to find peaks in the Hi-C read coverage with findPeaks.
One important note: the tagInfo.txt file will report the "Total Tags" value as 2x the total number interactions. This is because HOMER is storing each PE tag twice - the software later adjusts this number to reflect this redundancy, but keep this in mind when interpreting this number.
To create a paired-end tag directory, you must run makeTagDirectory by entering the two halves of the mapped reads separated by a comma, without spaces. For example:
makeTagDirectory <OutputTagDirectory>
<alignment file-read1>,<alignment
file-read2> ... [-illuminaPE] [-tbp 1]
i.e. makeTagDirectory ES-HiC lane1-read1.sam,lane1-read2.sam lane2-read1.sam,lane2-read2.sam -illuminaPE -tbp 1
i.e. makeTagDirectory ES-HiC lane1-read1.sam,lane1-read2.sam lane2-read1.sam,lane2-read2.sam -illuminaPE -tbp 1
HOMER matches reads between alignment files by matching their names, but to work with the original bowtie output files for Illumina sequencing (which often contains a 0 or 1 at the end to delineate the read source), it will need to remove the last character before trying to match. In this case, add "-illuminaPE" to the end of the command. If you download reads from the SRA they often have the same name for each paired end read - in these cases you don't want to remove the last character, so do NOT add "-illuminaPE" to the end of the command in that case.
Another typical option to add for Hi-C data is "-tbp 1", which will only consider read pairs with the exact same ends once. Given the search space in Hi-C, you would never expect to see reads-pairs start from the exact same places twice in the data (at least in the case of sonication). If you do, they are likely clonal and a result of over-sequencing your library or some other artifact.
On a side note, the HOMER tag directory created for paired-end reads is very similar to the directories created when running makeTagDirectory for single-end reads. The paired end reads are stored in *.tags.tsv files (extended format to include the pair), only each read will get 2 entries in the appropriate *.tags.tsv so that the reads can be indexed on each chromosome in one file. A flag is also set in the tagInfo.txt file to inform HOMER to treat the experiment as a paired-end file. Many of the programs in HOMER, such as makeUCSCfile, will treat paired-end tag directories as "single-end" directories to create output. This can be useful if you want to visualize read coverage in the UCSC Genome Browser or want to find peaks in the Hi-C read coverage with findPeaks.
One important note: the tagInfo.txt file will report the "Total Tags" value as 2x the total number interactions. This is because HOMER is storing each PE tag twice - the software later adjusts this number to reflect this redundancy, but keep this in mind when interpreting this number.
Alternative Input Files
HOMER will also accept "Hi-C Summary" files (use "-format HiCsummary"), which is a simple text file format that contains the mapping positions for a read-pair on each line. These files are tab-delimited text files with the following column assignments:
Hi-C Summary Format (columns):An example of using a Hi-C summary file:
1. Read Name (can be blank)
2. chromosome for read 1
3. positions for read 1 (5' end of read, one-indexed)
4. strand of read 1 (+ or -)
5. chromosome for read 2
6. positions for read 2 (5' end of read, one-indexed)
7. strand of read 2 (+ or -)
makeTagDirectory proB-HiC -format HiCsummary proB.HiC.FA.summary.txtYou can also combine Hi-C tag directories or add *tags.tsv files, just like with the regular makeTagDirectory command. For example, to combine directories:
makeTagDirectory ES-HiC-All -d ES-HiC-rep1/ ES-HiC-rep2/ ES-HiC-rep3/
Hi-C Quality Control
When running the makeTagDirectory
program, several standard quality control analysis files
will be created that resemble the analysis carried out for
single-end reads. You can also check for GC bias by
specifying the genome (i.e. "-genome hg19") and "-checkGC" at the
command line (analysis will be conducted by treating the
samples as "single" reads).
The following two Hi-C/paired-end read specific QC files will also be produced by default:
petagLocalDistribution.txt
The following two Hi-C/paired-end read specific QC files will also be produced by default:
petagLocalDistribution.txt
This file is similar to
the autocorrelation file, except that it shows the
relationship between the 5' ends of the paired
reads. This data is used to determine the fragment
size of the Hi-C fragments used for sequencing, assuming
that we should see a natural peak from "re-ligation
events" or random genomic fragments that got into the
sequencing library. Below is an example
(Here we'd estimate the fragment length to be about 350
bp):
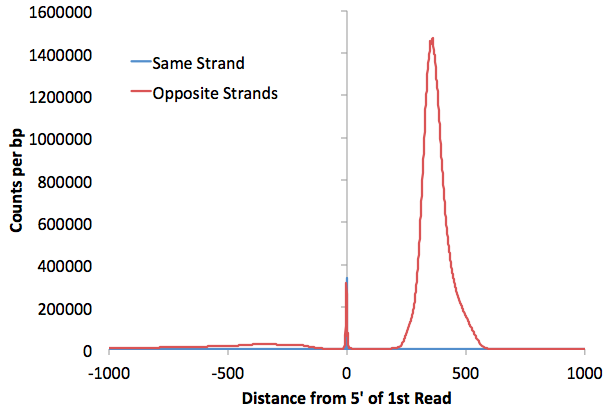
petagDistDistribution.txt
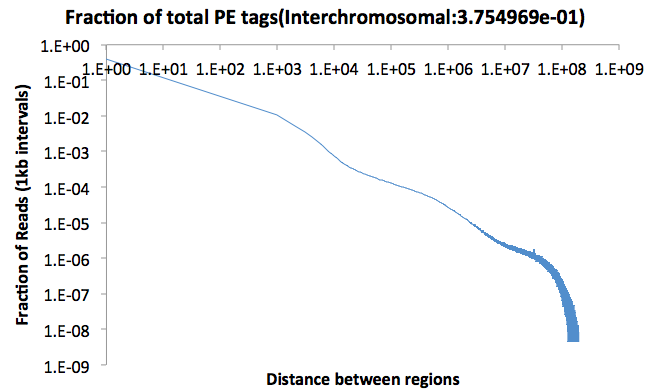
This file contains a
histogram describing the fraction of paired-end reads
that are found at different distances from one
another. The file stops at 300 million bp, and is
divided into 1kb bins. The top of the file also
contains the fraction of interchromosomal
interactions. Below is an example (plotted as a
log vs. log plot). The fraction of PE reads with
interactions greater than 300 Mb in distance are shown
in the final point.

Restriction Sites
One major problem that can
arise during Hi-C is that "background" ligation events may
occur. These are ligation events that occur after
sonication [I think] as they seem to occur between random
fragments of DNA with no regard to restriction site used
for the Hi-C assay. In theory, all reads should be
in the vicinity of the restriction site used for the
assay, and if they aren't, there is a good chance they are
noise (more later on this). To see the distribution
of reads around your restriction site, use the option "-restrictionSite <seq>"
(i.e. "-restrictionSite
AAGCTT"). You MUST specify the genome so
that HOMER can figure out where the restriction sites are
located (i.e. "-genome
mm9"). If the restriction site has a lot of
star activity, you can also add "-rsmis <#>" (i.e. "-rsmis 1") to indicate
how many mismatches are tolerated in the sites. By
default it only looks for perfect restriction sites.
Adding these options will produce a file named "petagRestrictionDistribution.<seq>.mis<#>.txt" (such as petagRestrictionDistribution.AAGCTT.mis0.txt). This file will display the distribution of reads relative to the restriction site. Graphing it with Excel will look something like this:
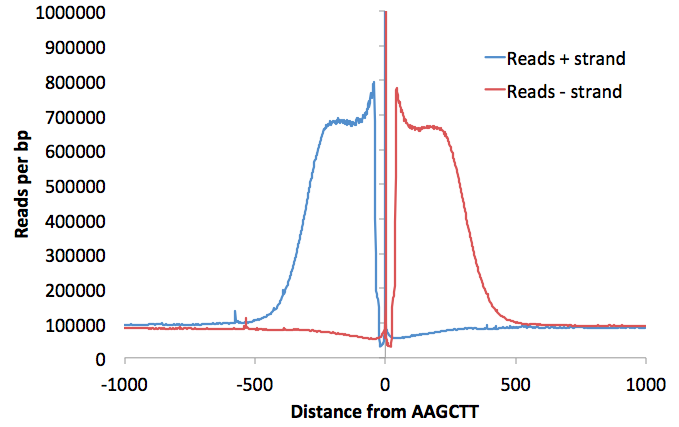
Adding these options will produce a file named "petagRestrictionDistribution.<seq>.mis<#>.txt" (such as petagRestrictionDistribution.AAGCTT.mis0.txt). This file will display the distribution of reads relative to the restriction site. Graphing it with Excel will look something like this:

Filtering Uninformative Reads
The Hi-C protocol can
produce sequencing libraries that contain many read pairs
that are not representative of true interactions.
This includes continuous genomic fragments, self-ligation
or re-ligation products, and background non-specific
ligation that occurs independent of a restriction site.
-removePEbg : This will remove paired-end reads that are likely continuous genomic fragments or re-ligation events. If this option is specified, read pairs are removed if they are separated less than 1.5x the sequencing insert fragment length. This distance is estimated from the distribution found in the file "petagLocalDistribution.txt" described above. If you wish to specify your own size, use "-fragLength <#>" to set this length manually.
-removeSpikes <#size> <# fold> : This option will remove PE reads that originate from regions of unusually high tag density. These regions arise for various reasons, for example, a region was duplicated in a given cell line. These types of anomalies tend to create artifacts in the data. Removing these from the analysis can help clean things up. The size parameter is the window of tag density that will be examined to find anomalies, and #-fold is how many tags over the average is needed to determine if a locus should be removed from the dataset. I'd recommend something like "-removeSpikes 10000 5" to get ride of the regions that are really bad. For example, this will scan all 10kb regions in the genome, calculate the average read density, and remove reads from regions that contain more than 5x the average number of reads.
-removePEbg : This will remove paired-end reads that are likely continuous genomic fragments or re-ligation events. If this option is specified, read pairs are removed if they are separated less than 1.5x the sequencing insert fragment length. This distance is estimated from the distribution found in the file "petagLocalDistribution.txt" described above. If you wish to specify your own size, use "-fragLength <#>" to set this length manually.
-removeSpikes <#size> <# fold> : This option will remove PE reads that originate from regions of unusually high tag density. These regions arise for various reasons, for example, a region was duplicated in a given cell line. These types of anomalies tend to create artifacts in the data. Removing these from the analysis can help clean things up. The size parameter is the window of tag density that will be examined to find anomalies, and #-fold is how many tags over the average is needed to determine if a locus should be removed from the dataset. I'd recommend something like "-removeSpikes 10000 5" to get ride of the regions that are really bad. For example, this will scan all 10kb regions in the genome, calculate the average read density, and remove reads from regions that contain more than 5x the average number of reads.
Filtering Reads based on Restriction Sites
All the options in this
section depend on restriction sites and require the
following to be part of the command:
-restrictionSite <seq> -genome <genome>
Restriction Site Proximity: The following options can be used to specify which reads should be kept with respect to the location of nearby restriction sites. The distance used to assign reads to restriction site is 1.5x the fragment length by default (set it with -restrictionSiteLength <#>):
-removeSelfLigation : Remove reads if their ends form a self ligation with adjacent restriction sites.
-removeRestrictionEnds : Removes reads if one of their ends starts on a restriction site.
-restrictionSiteLength <#> : Maximum distance to a restriction site for assignment.
-restrictionSite <seq> -genome <genome>
Restriction Site Proximity: The following options can be used to specify which reads should be kept with respect to the location of nearby restriction sites. The distance used to assign reads to restriction site is 1.5x the fragment length by default (set it with -restrictionSiteLength <#>):
-both : Only keep reads if both ends of the paired-end read have a restriction site within the fragment length estimate 3' to the read. This is the most conservative, and probably recommended for most types of analysis.
-one : Only keep reads if one or both ends of the paired-end read are near restriction sites.
-onlyOne : Only keep reads if one (and only one) of the reads in a paired-end read are near a restriction sites (mainly used for troubleshooting).
-none : Only keep read if neither of the ends are near a restriction site (mainly used for troubleshooting)
-removeSelfLigation : Remove reads if their ends form a self ligation with adjacent restriction sites.
-removeRestrictionEnds : Removes reads if one of their ends starts on a restriction site.
-restrictionSiteLength <#> : Maximum distance to a restriction site for assignment.
Recommended Settings:
The following is an example of how an average experiment should probably be analyzed. In this example we'll make an initial tag directory and remove all clonal reads, then make a copy of the directory, and then filter reads based on HindIII restriction sites:
makeTagDirectory ES-HiC-unfiltered reads1-1.sam,reads1-2.sam -tbp 1
cp -r ES-HiC-unfiltered ES-HiC-filtered
makeTagDirectory ES-HiC-filtered -update -genome hg19 -removePEbg -restrictionSite AAGCTT -both -removeSelfLigation -removeSpikes 10000 5
Command Line Options:
Usage: makeTagDirectory <directory> <alignment file 1> [file 2] ... [options]
Creates a platform-independent 'tag directory' for later analysis.
Currently BED, eland, bowtie, and sam files are accepted. The program will try to
automatically detect the alignment format if not specified. Program will also
unzip *.gz, *.bz2, and *.zip files and convert *.bam to sam files on the fly
Existing tag directories can be added or combined to make a new one using -d/-t
If more than one format is needed and the program cannot auto-detect it properly,
make separate tag directories by running the program separately, then combine them.
To perform QC/manipulations on an existing tag directory, add "-update"
Options:
-fragLength <# | given> (Set estimated fragment length - given: use read lengths)
By default treats the sample as a single read ChIP-Seq experiment
-format <X> where X can be: (with column specifications underneath)
bed - BED format files:
(1:chr,2:start,3:end,4:+/- or read name,5:# tags,6:+/-)
-force5th (5th column of BED file contains # of reads mapping to position)
sam - SAM formatted files (use samTools to covert BAMs into SAM if you have BAM)
-unique (keep if there is a single best alignment based on mapq)
-mapq <#> (Minimum mapq for -unique, default: 10, set negative to use AS:i:/XS:i:)
-keepOne (keep one of the best alignments even if others exist)
-keepAll (include all alignments in SAM file)
-mis (Maximum allowed mismatches, default: no limit, uses MD:Z: tag)
bowtie - output from bowtie (run with --best -k 2 options)
(1:read name,2:+/-,3:chr,4:position,5:seq,6:quality,7:NA,8:misInfo)
eland_result - output from basic eland
(1:read name,2:seq,3:code,4:#zeroMM,5:#oneMM,6:#twoMM,7:chr,
8:position,9:F/R,10-:mismatches
eland_export - output from illumina pipeline (22 columns total)
(1-5:read name info,9:sequence,10:quality,11:chr,13:position,14:strand)
eland_extended - output from illumina pipeline (4 columns total)
(1:read name,2:sequence,3:match stats,4:positions[,])
mCpGbed - encode style mCpG reporting in extended BED format, no auto-detect
(1:chr,2:start,3:end,4:name,5:,6:+/-,7:,8:,9:,10:#C,11:#mC)
-minCounts <#> (minimum number of reads to report mC/C ratios, default: 10)
HiCsummary - minimal paired-end read mapping information
(1:readname,2:chr1,3:5'pos1,4:strand1,5:chr2,6:5'pos2,7:strand2)
-force5th (5th column of BED file contains # of reads mapping to position)
-d <tag directory> [tag directory 2] ... (add Tag directory to new tag directory)
-t <tag file> [tag file 2] ... (add tag file i.e. *.tags.tsv to new tag directory)
-single (Create a single tags.tsv file for all "chromosomes" - i.e. if >100 chromosomes)
-update (Use current tag directory for QC/processing, do not parse new alignment files)
-tbp <#> (Maximum tags per bp, default: no maximum)
-precision <1|2|3> (number of decimal places to use for tag totals, default: 1)
GC-bias options:
-genome <genome version> (To see available genomes, use "-genome list")
-or- (for custom genomes):
-genome <path-to-FASTA file or directory of FASTA files>
-checkGC (check Sequence bias, requires "-genome")
-freqStart <#> (offset to start calculating frequency, default: -50)
-freqEnd <#> (distance past fragment length to calculate frequency, default: +50)
-oligoStart <#> (oligo bias start)
-oligoEnd <#> (oligo bias end)
-normGC <target GC profile file> (i.e. tagGCcontent.txt file from control experiment)
Use "-normGC default" to match the genomic GC distribution
-normFixedOligo <oligoFreqFile> (normalize 5' end bias, "-normFixedOligo default" ok)
-minNormRatio <#> (Minimum deflation ratio of tag counts, default: 0.25)
-maxNormRatio <#> (Maximum inflation ratio of tag counts, default: 2.0)
Paired-end/HiC options
-illuminaPE (when matching PE reads, assumes last character of read name is 0 or 1)
-removePEbg (remove paired end tags within 1.5x fragment length on same chr)
-PEbgLength <#> (remove PE reads facing on another within this distance, default: 1.5x fragLen)
-restrictionSite <seq> (i.e. AAGCTT for HindIII, assign data < 1.5x fragment length to sites)
Must specify genome sequence directory too. (-rsmis <#> to specify mismatches, def: 0)
-both, -one, -onlyOne, -none (Keeps reads near restriction sites, default: keep all)
-removeSelfLigation (removes reads linking same restriction fragment)
-removeRestrictionEnds (removes reads starting on a restriction fragment)
-assignMidPoint (will place reads in the middle of HindIII fragments)
-restrictionSiteLength <#> (maximum distance from restriction site, default: 1.5x fragLen)
-removeSpikes <size bp> <#> (remove tags from regions with > than # times
the average tags per size bp, suggest "-removeSpikes 10000 5")

Can't figure something out? Questions, comments, concerns, or other feedback:
cbenner@ucsd.edu