HOMER
Software for motif discovery and next-sequencing analysis
Selecting Background Sequences (homer2 background)
Traditionally, HOMER has selected background
sequences semi-randomly from the genome, making sure that
the per-region GC% distribution in the background sequences
matched that of the target sequences, explained here. HOMER2
now offers much greater control of the background sequence,
including routines that help model position-specific
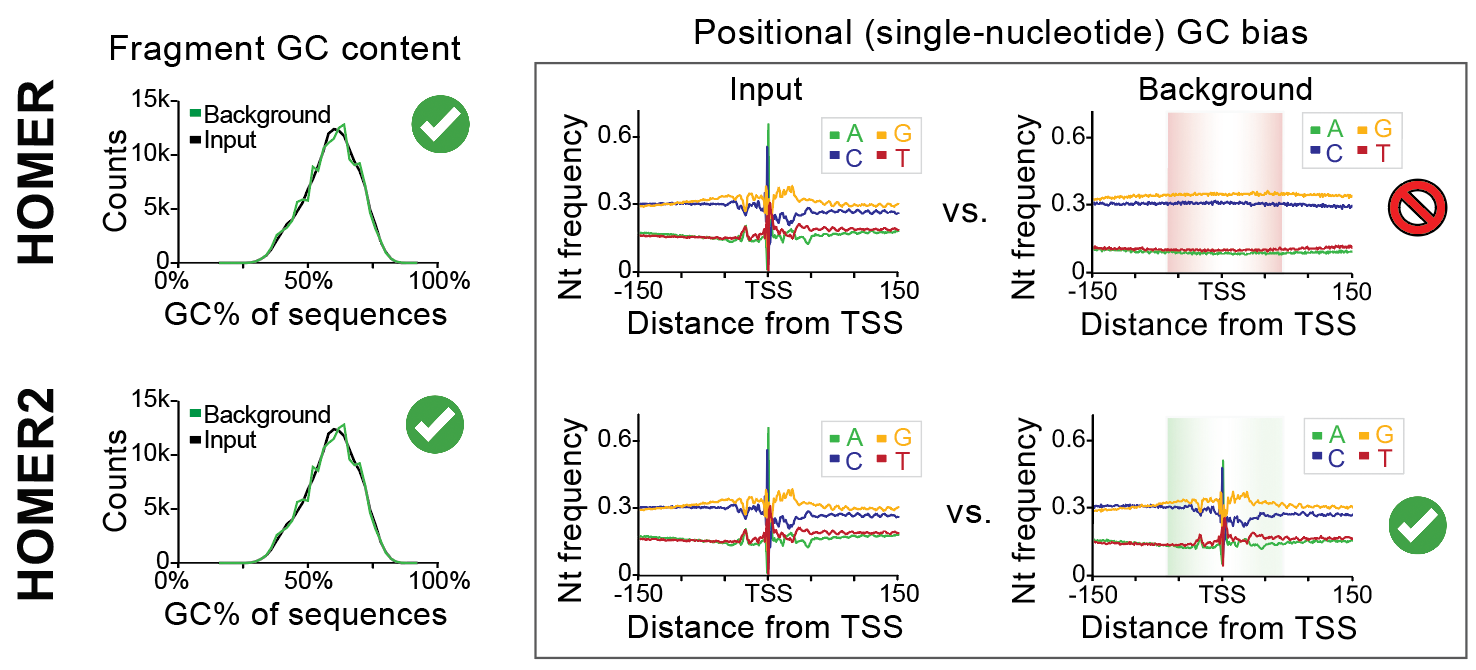
features of background sequence.The new background features are performed using a new subprogram called "homer2 background". This tool allows HOMER to randomly select from the genome or generate synthetic sequences that match positional k-mer content. This means that if your DNA sequences are anchored on a feature of interest, HOMER will attempt to account for lower-order (i.e. dinucleotide) sequence bias that might be present in your sequences. For example, if you have sequences centered on transcription start sites (TSS), certain nucleotides are likely to occur at certain positions, which may artificially increase the chance a given DNA motif will be recognized at that position. New background selection methods help address this concern, and provide tools to score the relative enrichment/depletion of motifs as a function of the motifs position in such sequences.
The "homer2 background" program is used by two other utilities, findMotifsGenome.pl and createHomer2EnrichmentTable.pl. However, if you want full control over how HOMER selects background sequences, you can use this tool directly. Furthermore, if you would like to use HOMER to generate background sequences for other applications or for use with other tools and analysis, you may want to use "homer2 background" directly. Ultimately, the output sequences can then be use as controls for DNA motif finding, enrichment calculations, machine learning applications, or other creative uses.
One important note that applies to all of these
approaches - the sequences you analyze must all have the SAME
LENGTH. The assumption for positional
enrichment is that all of your sequences are anchored on
some position or feature of interest such that there is a
relationship between nucleotide 1, 2, 3, etc. across all
of the input sequences.
homer2 background
The basic idea is to select background sequences with similar position-specific sequence properties relative to your input/target sequences. Traditionally, HOMER only normalizes for GC content on a per-sequence basis, which helps control for the presence of sequences with extreme levels of GC nucleotides which are typically found in CpG Islands near vertebrate regulatory regions (e.g. promoters). With HOMER2, not only will homer control for the overall per-sequence GC%, it will also attempt to select (or model sequences) that contain similar lower-order nucleotide content (i.e. mononuclotides, dinucleotides, trinucleotides, etc.) at each position.One thing to keep in mind is that this program was designed with regulatory elements in mind. It is meant to work with collections of sequences that are ~50 to 500 bp in length comprised of ~5000 to 500,000 total sequences (although it can work in other scenarios too).
Key inputs and parameters:
- Input sequences (either genomic positions or a FASTA
file, MUST be the same length)
- Genome or other FASTA file to select background sequences from (technically optional as HOMER2 can create synthetic background sequences if desired).
- k = length of k-mer at each position to match sequence properties with (k=1 nucleotides, k=2 dinucleotides, k=3 trinucleotides, etc.)
- Position-dependent or position-independent analysis of k-mer content (i.e. normalize k-mers regardless of position)
- Select actual background sequences or generate
synthetic sequences.
HOMER2 then performs the following steps:
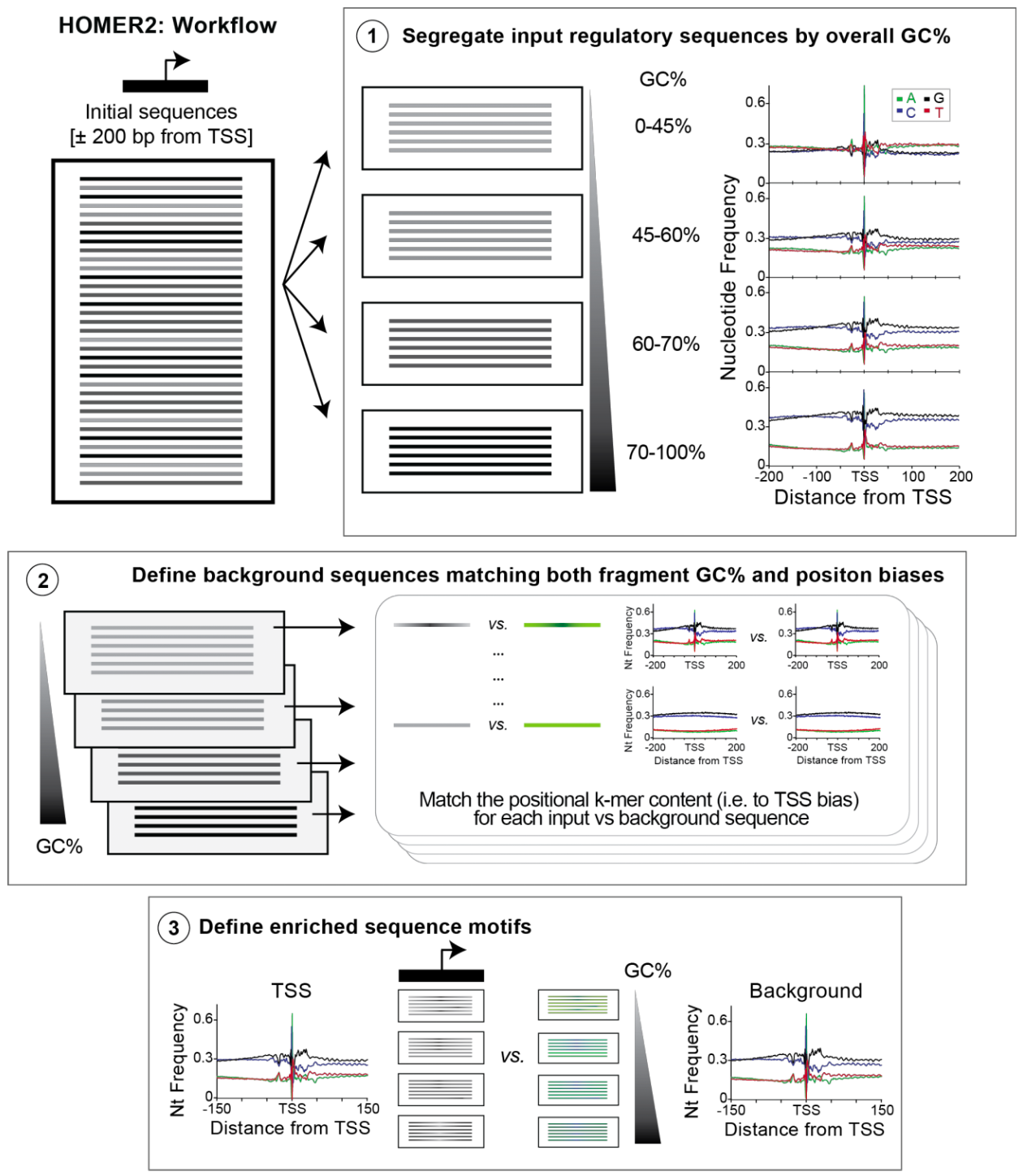
Sample Data:
- tss.positions.200.txt
- HOMER-style peak file similar to BED that contains
regions centered on 155k TSS locations identified by
csRNA-seq in U2OS cells in human [hg38]). The "200"
refers to the fact that the regions are 200 bp long
(centered on the TSS). Unzip it with gunzip before using
it.
- hg38.fa - you can download this directly from UCSC using "wget https://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/hg38.fa.gz" and then unzipping the file with gunzip
Example: This program will take ~5min and use ~16Gb of memory (if that is too much, try setting -NN 10000000 to only use ~8Gb):
homer2 bg -p tss.positions.200.txt -g hg38.fa -pkmer 2 -N 100000 -NN 100000000 -o output -allowTargetOverlap -allowBgOverlap
Output: The program will create several output files starting with <prefix> (set with -o <prefix>, in the case above it is 'output'):
- <prefix>.info.txt - reports the command line options used and some basic information including actual number of sequences in output background set and the cumulative 'difference' or error in the background sequence properties and the target set.
- <prefix>.bg.positions.bed - tab-delimited
BED format file with the genomic positions of all
selected sequences
- <prefix>.bg.sequences.fasta - FASTA
formatted file containing the selected background
sequences
- <prefix>.bg.stats.txt - table containing the name of the background sequence, chromosome, position (min regardless of strand), strand, weight (1.0, not used), Profile Score (internal score, not used), GC%, and the internal bin ID the background was assigned from.
- <prefix>.bg.positions.bed, <prefix>.bg.sequences.fasta,
and <prefix>.target.stats.txt provide the
same information, but for the target sequences.
The output sequences can then be use as controls for DNA motif finding, enrichment calculations, or other creative uses.
Note - there is a LOT more that this program can do to control how your background sequences are created. In particular, you can also use it to model potential background sequences instead of having them selected from the genome. See the background page for more information.
Usage:
homer2 background -i <target sequences.fasta> [options]
Generate/Select background sequences that match properties in a set of target sequences.
Inputs:
Target sequences you want to model:
-i <target sequences.fasta> (FASTA file)
-p <target positions.bed> (Alteratively, provide a BED or HOMER peak file with genomic coordinates)
Background sequences to select from:
-model (generate sequences using a model, do not extract real background sequences)
-g <genome.fasta> (genome FASTA file or seqeunce resource to select sequences from)
-b <background sequences.fasta> (explicit set of background sequences to choose from FASTA file)
-bg <background positions.bed> (explicit set of background positions to choose from)
-bgr <background regions positions.bed> (regions of the genome to select bg sequences from)
Key options:
-size <#> (size of regions to consider in background, default: avg of length of target sequences)
-N <#> (number of background sequences to select, default: 100000)
-NN <#> (number of background sequences initial screen from genome, default: 100000000)
-mask (mask lowercase sequence i.e. softmasked sequence, default: use all sequences)
-nbins <#> (number of bins to segregate sequences into for GC selection, def: 10
-nsubBins <#> (number of bins to segregate sequences into for positional frequencies, def: 10
-maxFractionN <#> (Maximum fraction of sequence that can be N and still used, default: 0.5)
-allowTargetOverlaps (allow selected bg sequences from a genome to overlap targets, def: not allowed)
-allowBgOverlaps (allow selected bg sequences from a genome to overlap, def: not allowed)
-strand (allow sequences to overlap if on separate strands)
-pkmer <#> (match positional kmer content)
-ikmer <#> (match overall kmer content [position independent])
-excludeNs/-includeNs (by default, kmers with Ns are excluded when selecting bg sequences,
but included when generating sequences with -model)
-pscore <outputBEDfile> (Report initial pscores)
-maxIterations <#> (maximum iterations, def: 20)
-overlapIteration <#> (iteration to start enforcing no overlaps, def: 5)
-decayRate <#> (selection rate per iteration, def: 0.75)
-seed <#> (seed for random number generator, def: uses time)
Output:
-o <output prefix> (default: out)
-gs (include homer-style group and sequence output files)
Important: You input sequences must be the same length!
Below are some general tips for getting the most out of you motif analysis when using HOMER. Be sure to look over this section about judging motif quality!
Why is the number of background regions reported by HOMER different then my input files?
HOMER performs a step to normalize the GC-content of the background sequences, which may result in the adjustment of the total apparent number of background sequences. If you target sequences are GC-rich and your background sequences are AT-rich (a common issue with mammalian genomes), many of the AT-rich sequences may be added fractionally to the total so that the imbalance is minimized.
Why do motif counts from findMotifsGenome.pl and annotatePeaks.pl differ?
By default, annotatePeaks.pl uses the given size of the peaks (default: -size given), while findMotifsGenome.pl uses a default size of 200 (default: -size 200). NO

Can't figure something out? Questions, comments, concerns, or other feedback:
cbenner@ucsd.edu